You will later see that we use this operator more often, e.g., when we create moves and monitors. This simply means that the variable is not part of the model. You may have noticed that we used the = operator to create the move index. To check your current working directory, use the function getwd().

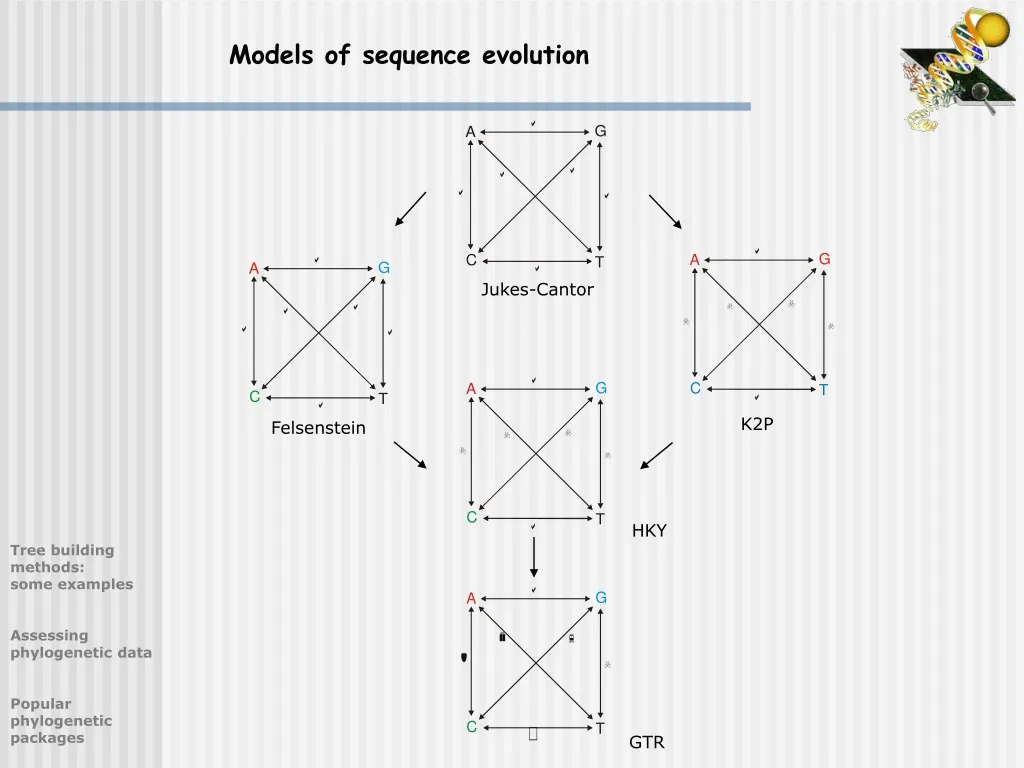
In your Rev scripts or in the RevBayes console. It is important that you are aware of your current working directory if you use relative file paths Checking and Changing Your Working Directoryįor this tutorial and much of the work you will do in RevBayes, you will need to access files. Now start RevBayes from your working directory ( RB_CTMC_Tutorial). Navigate to this new directory and create a new folder called data inside of it.ĭownload the data file called primates_and_galeopterus_cytb.nex Loading the Dataįirst create a directory for this tutorial and name it RB_CTMC_Tutorial, or any name The next steps will walk you through the full specification of the model and MCMC analyses. Furthermore, without inspecting the Rev scripts sourced in mcmc_JC.Rev, you may end up inadvertently performing inappropriate analyses on your dataset, which would be a waste of your time and CPU cycles. However, it is important to understand the components of the model to be able to take full advantage of the flexibility and richness of RevBayes. You could easily run this entire analysis on your own data by substituting your data file name for that in the model-specification file. Ultimately, this is how you will execute most analyses in RevBayes, with the full specification of the model and analyses contained in the sourced files. Output the states of the Markov chain once the MCMC analysis begins. If you continue to let this run, then you will see it The Markov chain Monte Carlo analysis that estimates the posteriorĭistribution. If everything loaded properly, then you should see the program initiate If you download this file and place it in a directory called scripts inside your main tutorial directory,Įasily execute this analysis using the source() function in the RevBayes console: The files for this example analysis are provided for you ( mcmc_JC.Rev). You will be able to complete the exercises without understanding the underlying math. Here we only provide some of the equations for the models in case you might be interested in the details. Don’t worry, you won’t have to calculate all of the transition probabilities, because RevBayes will take care of all the computations for you. In the later exercises you will be asked to specify more complex substitution models. Where $t$ is the branch length in units of time, and $r$ is the rate (clock) for the process. Used in molecular evolution are continuous time Markov models, which areįully characterized by their instantaneous-rate matrix:

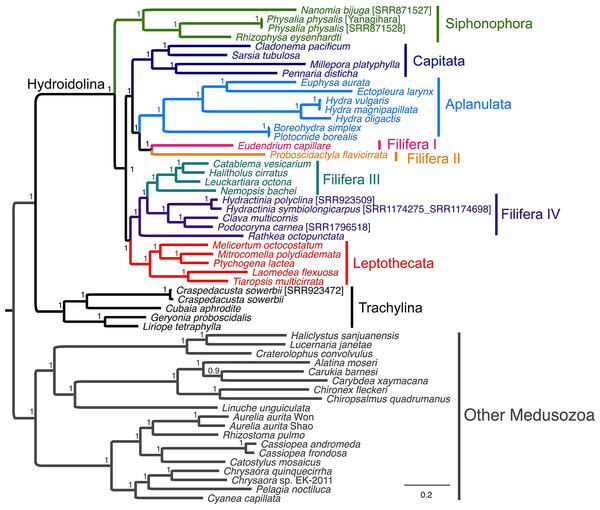
Using common nucleotide substitution models. Which demonstrates how to set up and perform analyses This tutorial covers the first protocol from Höhna et al. The full playlist is available here: Overview The video corresponding to each section of the exercise is linked next to the section title.
This tutorial comes with a recorded video walkthrough. Introduction to Markov chain Monte Carlo (MCMC) Sampling.Getting Started with RevBayes and Rev Language Syntax.
